Automated, distributed image processing pipelines
From Cloud Services to Service Pipelines
In its first years, HIFIS has set up the Helmholtz Cloud to interconnect and broker services from Helmholtz for Helmholtz. The Helmholtz Cloud Portal is the central point where users have access to all services and combine them to suit their work flow.
Interconnecting cloud services — sometimes dubbed Service Orchestration — allows to automate complex workflows, a precondition to ultimately set up reliable, scalable, reproducible and adaptable data processing pipelines for science. A typical example is to automatically and continuously fetch, move, process, publish and store scientific data, possibly in multiple iterations and for different audiences.
Such interconnection of services and resources can in principle be implemented by end users, thanks to the open interfaces of most offered cloud services and the service-agnostic authentication and authorisation infrastructure (AAI) with its single-sign on functionality.
However: This is the point where extensive knowledge on the underlying technologies is necessary in order to implement complex service orchestration workflows.
Whenever you plan to implement such complex use cases yourself: Yay!! Feel free to contact us — We are always happy to hear from you and even happier to assist!
Mostly, though, you will probably be glad to use (and adopt) pre-confectioned pipelines. So, let’s see what’s already possible!

A Pilot Use Case from Helmholtz Imaging
Helmholtz Imaging, a ‘‘sister’’ platform to HIFIS, aims to improve access for scientists on innovative scientific imaging modalities, methodological richness, and data treasures within the Helmholtz Association. HIFIS can support this mission through seamless interconnection of the cutting-edge IT infrastructures in Helmholtz as orchestrated services.
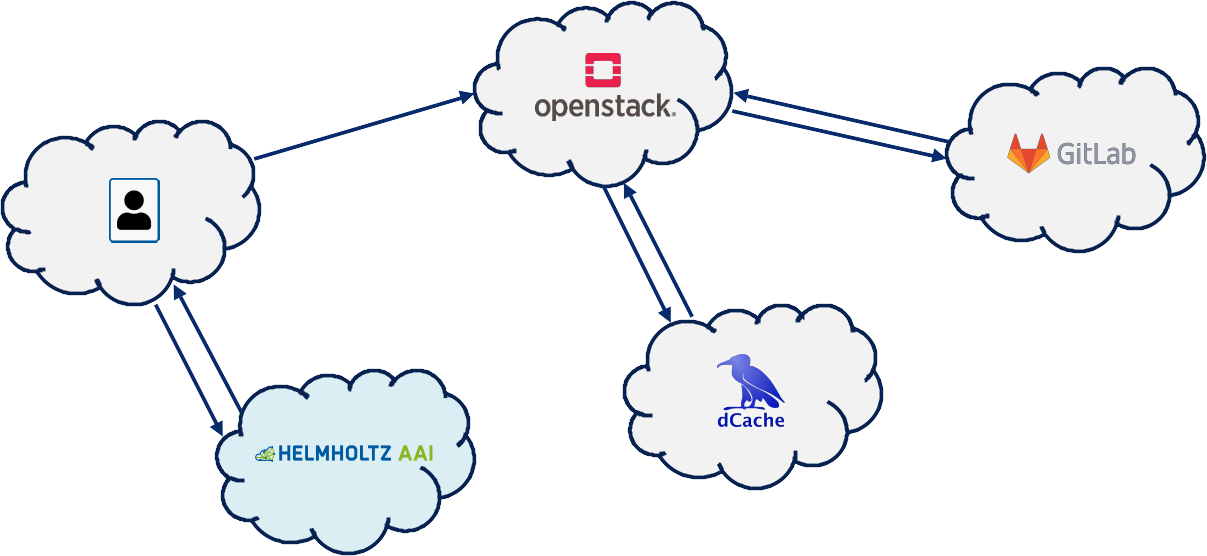
Our pilot implementation of such orchestration showcases the interconnectivity of five different Helmholtz centres: Cloud services are located in three centres, an interconnecting component is developed by a fourth, and the scientific user comes from a fifth centre.
The pilot also demonstrates the modularity of such pipelines, allowing to exchange or scale up parts of the pipeline, whenever beneficial.
The data we use to showcase our pipeline originates from a research paper published in 2020 [1]. The authors looked into the role of intracellular organelles, such as microtubules, for insulin transportation and secretion. They segmented organelles from high-resolution tomographic images, and provided mesh generation and rendering of 3D organelle reconstructions in primary mouse betacells as a model system.

Workflow in a nutshell
Using this exemplary data, we show how the data is handled and processed, starting from raw images until ‘‘publication’’. For more details, you may also check our presentation on an earlier version of this demonstrator .
- The primary data of the researcher (HZB) was stored in a Sync&Share. We do not advise to do this, as there are more sophisticated data storage solutions such as dCache InfiniteSpace (DESY). So, we transfer this first data to the dCache for all later usage and updates.
- The raw data, now located at DESY, is then transferred to a virtual machine located in Jülich, to do the processing there, with intermediate and final results sync’ed back to dCache at DESY, and prepared for publication on an open website built and hosted at HZDR.
- Almost the complete workflow is automated. The script magic is supplied via Helmholtz Codebase (Gitlab) at HZDR, allowing for collaborative, versioned development of scripts and further usage-specific magic.
- Minimal user interaction is necessary in the beginning for the primary login via AAI; no user interaction at all for subsequent iterations / updates of the data.
- The secure transfer of user login information between the services, without further manual interaction by the user, is mediated by a technology calls OIDC Agent, provided by KIT.


Why should I use this?
- All transfers and steps occur within Helmholtz infrastructure, with secure underlying authentication and authorisation methods. That is, all intermediate steps can be finely controlled to be private, group-shared and/or public, just as desired. Likewise, different steps of the pipeline can be set up to be accessible by different user (groups).
- The workflow can be set up to persist for a definable time frame, even when the original user(s) are not online anymore. For example, newly incoming raw data on dCache can trigger the pipeline to be continuously updated and provide correspondingly updated results.
- You need increased or reduced compute power at different points in time? You want to exchange the storage because the majority of primary data piles up somewhere else? You want to introduce another specific processing step employing another cloud service? No big deal: The modularity of the whole set-up allows to exchange or replace any part of the pipeline, whenever needed.
Use and combine the capabilities of Helmholtz Cloud to your best, without needing to solely rely on local resources.
That sounds cool. But… how to set this up?
The confection of this kind of orchestrated IT services and pipelines for specific classes of use cases will be a major focus of HIFIS from now on.
If you have a corresponding use case — any sort of complexity is welcome — please get in touch with us. We will be happy to assist and build cool stuff!
In the near future, generalized pipelines will be made available via our pages and the cloud portal, so: Stay tuned!
Get in contact
For HIFIS: HIFIS Support For Helmholtz Imaging: Helmholtz Imaging Helpdesk
Literature
Changelog
- 2023-06-28: Added Link to Helmholtz Cloud portal entry for newly onboarded dCache service, adapted wording
- 2025-05-19: Updated some links to Helmholtz Cloud Portal.